MEDTO: Medical Data to Ontology Matching using Hybrid Graph Neural Networks
Abstract
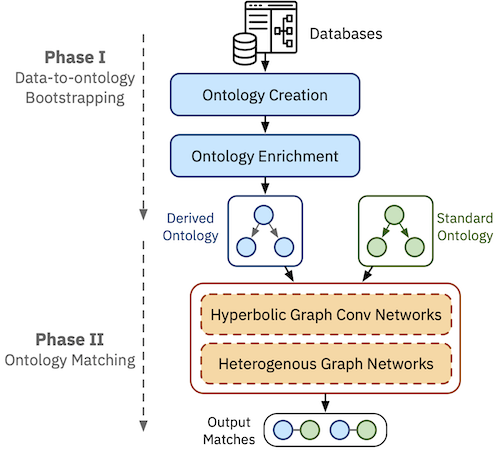
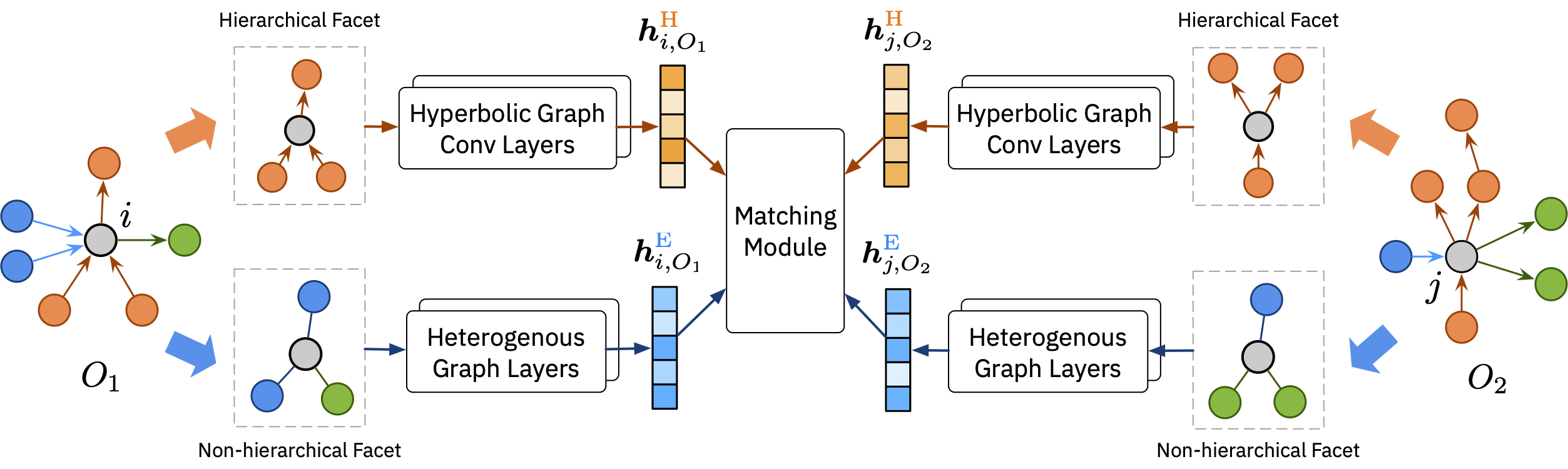
Medical ontologies are widely used to describe and organize medical terminologies and to support many critical applications on healthcare databases. These ontologies are often manually curated (e.g., UMLS, SNOMED CT, and MeSH) by medical experts. Medical databases, on the other hand, are often created by database administrators, using different terminology and structures. The discrepancies between medical ontologies and databases compromise interoperability between them. Data to ontology matching is the process of finding semantic correspondences between tables in databases to standard ontologies. Existing solutions such as ontology matching have mostly focused on engineering features from terminological, structural, and semantic model information extracted from the ontologies. However, this is often labor-intensive and the accuracy varies greatly across different ontologies. Worse yet, the ontology capturing a medical database is often not given in practice. In this paper, we propose MEDTO, a novel end-to-end framework that consists of three innovative techniques: (1) a lightweight yet effective method that bootstrap a semantically rich ontology from a given medical database, (2) a hyperbolic graph convolution layer that encodes hierarchical concepts in the hyperbolic space, and (3) a heterogeneous graph layer that encodes both local and global context information of a concept. Experiments on two real-world medical datasets matching against SNOMED CT show significant improvements compared to the state-of-the-art methods. MEDTO also consistently achieves competitive results on a benchmark from the Ontology Alignment Evaluation Initiative.
Modeling
More Info
Publication: The 27th International ACM SIGKDD Conference on Knowledge Discovery and Data Mining (KDD 2021)
Date: August, 2021
Links: Paper (ACM Library), PDF, Slides, Poster, Video